Mathematical Methods for Data Science
Keith Dillon
Spring 2020

Topic 2: "Classical" Linear Algebra Software
This topic:¶
- BLAS, LAPACK, Eigen
- Overview of major python packages
- Structured and Sparse Linear Algebra
- Numpy basics
Reading: the internet
What is "Classical" Linear Algebra software?¶
- A term I just invented
- Generally single CPU, RAM-limited implementations of well-known algorithms (e.g. Golub & Van Loan "Matrix Computations")
- ...and higher-level packages based on these algorithms, inheriting the limitations
- Large-scale (more than 10k variable or so) only possible via sparse linear algebra algorithms
- Leading parallel tool is SIMD (single instruction multiple data) via math co-processors.
- More parralelism at top-level rather than bottom ("workers" running seperate processes on separate CPU's)
- current growth is in rewriting basic algorithms for
I. BLAS BLAS BLAS¶
BLAS = Basic Linear Algebra Subroutines¶
Efficient algorithms for matrix algebra
- scaling, multiplying. "Level 1,2,3 functions"
- single and double precision, real and complex numbers
- generally don't take advantage of multiple threads or CPU's by chip maker
- Are often optimized for complex instructions however, e.g. SIMD
- Originally FORTRAN - one-based, columns first, like matlab
- various implementations, including parallelizable versions, C/C++ CUDA,
- http://www.netlib.org/blas/
BLAS Levels 1,2,3 ~ $O(n), O(n^2), O(n^3)$¶

import scipy.linalg.blas as blas
dir(blas)
['__all__', '__builtins__', '__cached__', '__doc__', '__file__', '__loader__', '__name__', '__package__', '__spec__', '_blas_alias', '_cblas', '_fblas', '_get_funcs', '_np', '_type_conv', '_type_score', 'absolute_import', 'caxpy', 'ccopy', 'cdotc', 'cdotu', 'cgbmv', 'cgemm', 'cgemv', 'cgerc', 'cgeru', 'chbmv', 'chemm', 'chemv', 'cher', 'cher2', 'cher2k', 'cherk', 'chpmv', 'chpr', 'chpr2', 'crotg', 'cscal', 'cspmv', 'cspr', 'csrot', 'csscal', 'cswap', 'csymm', 'csyr', 'csyr2k', 'csyrk', 'ctbmv', 'ctbsv', 'ctpmv', 'ctpsv', 'ctrmm', 'ctrmv', 'ctrsm', 'ctrsv', 'dasum', 'daxpy', 'dcopy', 'ddot', 'dgbmv', 'dgemm', 'dgemv', 'dger', 'division', 'dnrm2', 'drot', 'drotg', 'drotm', 'drotmg', 'dsbmv', 'dscal', 'dspmv', 'dspr', 'dspr2', 'dswap', 'dsymm', 'dsymv', 'dsyr', 'dsyr2', 'dsyr2k', 'dsyrk', 'dtbmv', 'dtbsv', 'dtpmv', 'dtpsv', 'dtrmm', 'dtrmv', 'dtrsm', 'dtrsv', 'dzasum', 'dznrm2', 'find_best_blas_type', 'get_blas_funcs', 'icamax', 'idamax', 'isamax', 'izamax', 'print_function', 'sasum', 'saxpy', 'scasum', 'scnrm2', 'scopy', 'sdot', 'sgbmv', 'sgemm', 'sgemv', 'sger', 'snrm2', 'srot', 'srotg', 'srotm', 'srotmg', 'ssbmv', 'sscal', 'sspmv', 'sspr', 'sspr2', 'sswap', 'ssymm', 'ssymv', 'ssyr', 'ssyr2', 'ssyr2k', 'ssyrk', 'stbmv', 'stbsv', 'stpmv', 'stpsv', 'strmm', 'strmv', 'strsm', 'strsv', 'zaxpy', 'zcopy', 'zdotc', 'zdotu', 'zdrot', 'zdscal', 'zgbmv', 'zgemm', 'zgemv', 'zgerc', 'zgeru', 'zhbmv', 'zhemm', 'zhemv', 'zher', 'zher2', 'zher2k', 'zherk', 'zhpmv', 'zhpr', 'zhpr2', 'zrotg', 'zscal', 'zspmv', 'zspr', 'zswap', 'zsymm', 'zsyr', 'zsyr2k', 'zsyrk', 'ztbmv', 'ztbsv', 'ztpmv', 'ztpsv', 'ztrmm', 'ztrmv', 'ztrsm', 'ztrsv']
LAPACK = Linear Algebra Package¶
- Fortran, Uses BLAS as backend (can choose different implementations)
- http://www.netlib.org/lapack/
- https://github.com/Reference-LAPACK/lapack
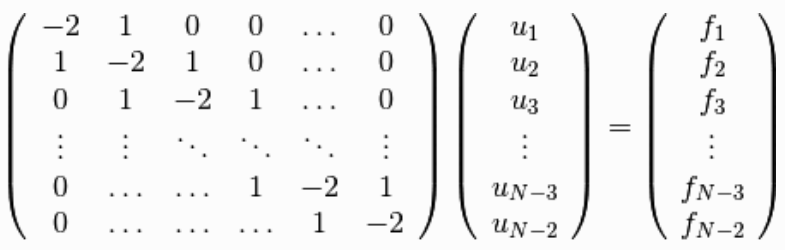
- systems of simultaneous linear equations
- least-squares solutions of linear systems of equations
- eigenvalue problems
- singular value problems.
- associated matrix factorizations (LU, Cholesky, QR, SVD, Schur, generalized Schur)
- related computations such as reordering of the Schur factorizations and estimating condition numbers.
- Dense and banded matrices are handled, but not general sparse matrices.
- real and complex matrices
- both single and double precision.
- Matlab, and the Python tools we will use were essentially built on this
import scipy.linalg.lapack as lapack
dir(lapack)
['_DeprecatedImport', '__all__', '__builtins__', '__cached__', '__doc__', '__file__', '__loader__', '__name__', '__package__', '__spec__', '_clapack', '_compute_lwork', '_dep_message', '_flapack', '_get_funcs', '_lapack_alias', '_np', 'absolute_import', 'cgbsv', 'cgbtrf', 'cgbtrs', 'cgebal', 'cgecon', 'cgees', 'cgeev', 'cgeev_lwork', 'cgegv', 'cgehrd', 'cgehrd_lwork', 'cgels', 'cgels_lwork', 'cgelsd', 'cgelsd_lwork', 'cgelss', 'cgelss_lwork', 'cgelsy', 'cgelsy_lwork', 'cgeqp3', 'cgeqrf', 'cgerqf', 'cgesdd', 'cgesdd_lwork', 'cgesv', 'cgesvd', 'cgesvd_lwork', 'cgesvx', 'cgetrf', 'cgetri', 'cgetri_lwork', 'cgetrs', 'cgges', 'cggev', 'cgglse', 'cgglse_lwork', 'cgtsv', 'chbevd', 'chbevx', 'checon', 'cheev', 'cheevd', 'cheevr', 'chegst', 'chegv', 'chegvd', 'chegvx', 'chesv', 'chesv_lwork', 'chesvx', 'chesvx_lwork', 'chetrd', 'chetrd_lwork', 'chetrf', 'chetrf_lwork', 'chfrk', 'clange', 'clapack', 'clarf', 'clarfg', 'clartg', 'claswp', 'clauum', 'cpbsv', 'cpbtrf', 'cpbtrs', 'cpftrf', 'cpftri', 'cpftrs', 'cpocon', 'cposv', 'cposvx', 'cpotrf', 'cpotri', 'cpotrs', 'cptsv', 'crot', 'csycon', 'csyconv', 'csysv', 'csysv_lwork', 'csysvx', 'csysvx_lwork', 'csytf2', 'csytrf', 'csytrf_lwork', 'ctfsm', 'ctfttp', 'ctfttr', 'ctgsen', 'ctpttf', 'ctpttr', 'ctrsyl', 'ctrtri', 'ctrtrs', 'ctrttf', 'ctrttp', 'ctzrzf', 'ctzrzf_lwork', 'cunghr', 'cunghr_lwork', 'cungqr', 'cungrq', 'cunmqr', 'cunmrz', 'cunmrz_lwork', 'dgbsv', 'dgbtrf', 'dgbtrs', 'dgebal', 'dgecon', 'dgees', 'dgeev', 'dgeev_lwork', 'dgegv', 'dgehrd', 'dgehrd_lwork', 'dgels', 'dgels_lwork', 'dgelsd', 'dgelsd_lwork', 'dgelss', 'dgelss_lwork', 'dgelsy', 'dgelsy_lwork', 'dgeqp3', 'dgeqrf', 'dgerqf', 'dgesdd', 'dgesdd_lwork', 'dgesv', 'dgesvd', 'dgesvd_lwork', 'dgesvx', 'dgetrf', 'dgetri', 'dgetri_lwork', 'dgetrs', 'dgges', 'dggev', 'dgglse', 'dgglse_lwork', 'dgtsv', 'division', 'dlamch', 'dlange', 'dlarf', 'dlarfg', 'dlartg', 'dlasd4', 'dlaswp', 'dlauum', 'dorghr', 'dorghr_lwork', 'dorgqr', 'dorgrq', 'dormqr', 'dormrz', 'dormrz_lwork', 'dpbsv', 'dpbtrf', 'dpbtrs', 'dpftrf', 'dpftri', 'dpftrs', 'dpocon', 'dposv', 'dposvx', 'dpotrf', 'dpotri', 'dpotrs', 'dptsv', 'dsbev', 'dsbevd', 'dsbevx', 'dsfrk', 'dstebz', 'dstein', 'dstemr', 'dstemr_lwork', 'dsterf', 'dstev', 'dsycon', 'dsyconv', 'dsyev', 'dsyevd', 'dsyevr', 'dsygst', 'dsygv', 'dsygvd', 'dsygvx', 'dsysv', 'dsysv_lwork', 'dsysvx', 'dsysvx_lwork', 'dsytf2', 'dsytrd', 'dsytrd_lwork', 'dsytrf', 'dsytrf_lwork', 'dtfsm', 'dtfttp', 'dtfttr', 'dtgsen', 'dtpttf', 'dtpttr', 'dtrsyl', 'dtrtri', 'dtrtrs', 'dtrttf', 'dtrttp', 'dtzrzf', 'dtzrzf_lwork', 'find_best_lapack_type', 'flapack', 'get_lapack_funcs', 'ilaver', 'print_function', 'sgbsv', 'sgbtrf', 'sgbtrs', 'sgebal', 'sgecon', 'sgees', 'sgeev', 'sgeev_lwork', 'sgegv', 'sgehrd', 'sgehrd_lwork', 'sgels', 'sgels_lwork', 'sgelsd', 'sgelsd_lwork', 'sgelss', 'sgelss_lwork', 'sgelsy', 'sgelsy_lwork', 'sgeqp3', 'sgeqrf', 'sgerqf', 'sgesdd', 'sgesdd_lwork', 'sgesv', 'sgesvd', 'sgesvd_lwork', 'sgesvx', 'sgetrf', 'sgetri', 'sgetri_lwork', 'sgetrs', 'sgges', 'sggev', 'sgglse', 'sgglse_lwork', 'sgtsv', 'slamch', 'slange', 'slarf', 'slarfg', 'slartg', 'slasd4', 'slaswp', 'slauum', 'sorghr', 'sorghr_lwork', 'sorgqr', 'sorgrq', 'sormqr', 'sormrz', 'sormrz_lwork', 'spbsv', 'spbtrf', 'spbtrs', 'spftrf', 'spftri', 'spftrs', 'spocon', 'sposv', 'sposvx', 'spotrf', 'spotri', 'spotrs', 'sptsv', 'ssbev', 'ssbevd', 'ssbevx', 'ssfrk', 'sstebz', 'sstein', 'sstemr', 'sstemr_lwork', 'ssterf', 'sstev', 'ssycon', 'ssyconv', 'ssyev', 'ssyevd', 'ssyevr', 'ssygst', 'ssygv', 'ssygvd', 'ssygvx', 'ssysv', 'ssysv_lwork', 'ssysvx', 'ssysvx_lwork', 'ssytf2', 'ssytrd', 'ssytrd_lwork', 'ssytrf', 'ssytrf_lwork', 'stfsm', 'stfttp', 'stfttr', 'stgsen', 'stpttf', 'stpttr', 'strsyl', 'strtri', 'strtrs', 'strttf', 'strttp', 'stzrzf', 'stzrzf_lwork', 'zgbsv', 'zgbtrf', 'zgbtrs', 'zgebal', 'zgecon', 'zgees', 'zgeev', 'zgeev_lwork', 'zgegv', 'zgehrd', 'zgehrd_lwork', 'zgels', 'zgels_lwork', 'zgelsd', 'zgelsd_lwork', 'zgelss', 'zgelss_lwork', 'zgelsy', 'zgelsy_lwork', 'zgeqp3', 'zgeqrf', 'zgerqf', 'zgesdd', 'zgesdd_lwork', 'zgesv', 'zgesvd', 'zgesvd_lwork', 'zgesvx', 'zgetrf', 'zgetri', 'zgetri_lwork', 'zgetrs', 'zgges', 'zggev', 'zgglse', 'zgglse_lwork', 'zgtsv', 'zhbevd', 'zhbevx', 'zhecon', 'zheev', 'zheevd', 'zheevr', 'zhegst', 'zhegv', 'zhegvd', 'zhegvx', 'zhesv', 'zhesv_lwork', 'zhesvx', 'zhesvx_lwork', 'zhetrd', 'zhetrd_lwork', 'zhetrf', 'zhetrf_lwork', 'zhfrk', 'zlange', 'zlarf', 'zlarfg', 'zlartg', 'zlaswp', 'zlauum', 'zpbsv', 'zpbtrf', 'zpbtrs', 'zpftrf', 'zpftri', 'zpftrs', 'zpocon', 'zposv', 'zposvx', 'zpotrf', 'zpotri', 'zpotrs', 'zptsv', 'zrot', 'zsycon', 'zsyconv', 'zsysv', 'zsysv_lwork', 'zsysvx', 'zsysvx_lwork', 'zsytf2', 'zsytrf', 'zsytrf_lwork', 'ztfsm', 'ztfttp', 'ztfttr', 'ztgsen', 'ztpttf', 'ztpttr', 'ztrsyl', 'ztrtri', 'ztrtrs', 'ztrttf', 'ztrttp', 'ztzrzf', 'ztzrzf_lwork', 'zunghr', 'zunghr_lwork', 'zungqr', 'zungrq', 'zunmqr', 'zunmrz', 'zunmrz_lwork']
C++ API¶
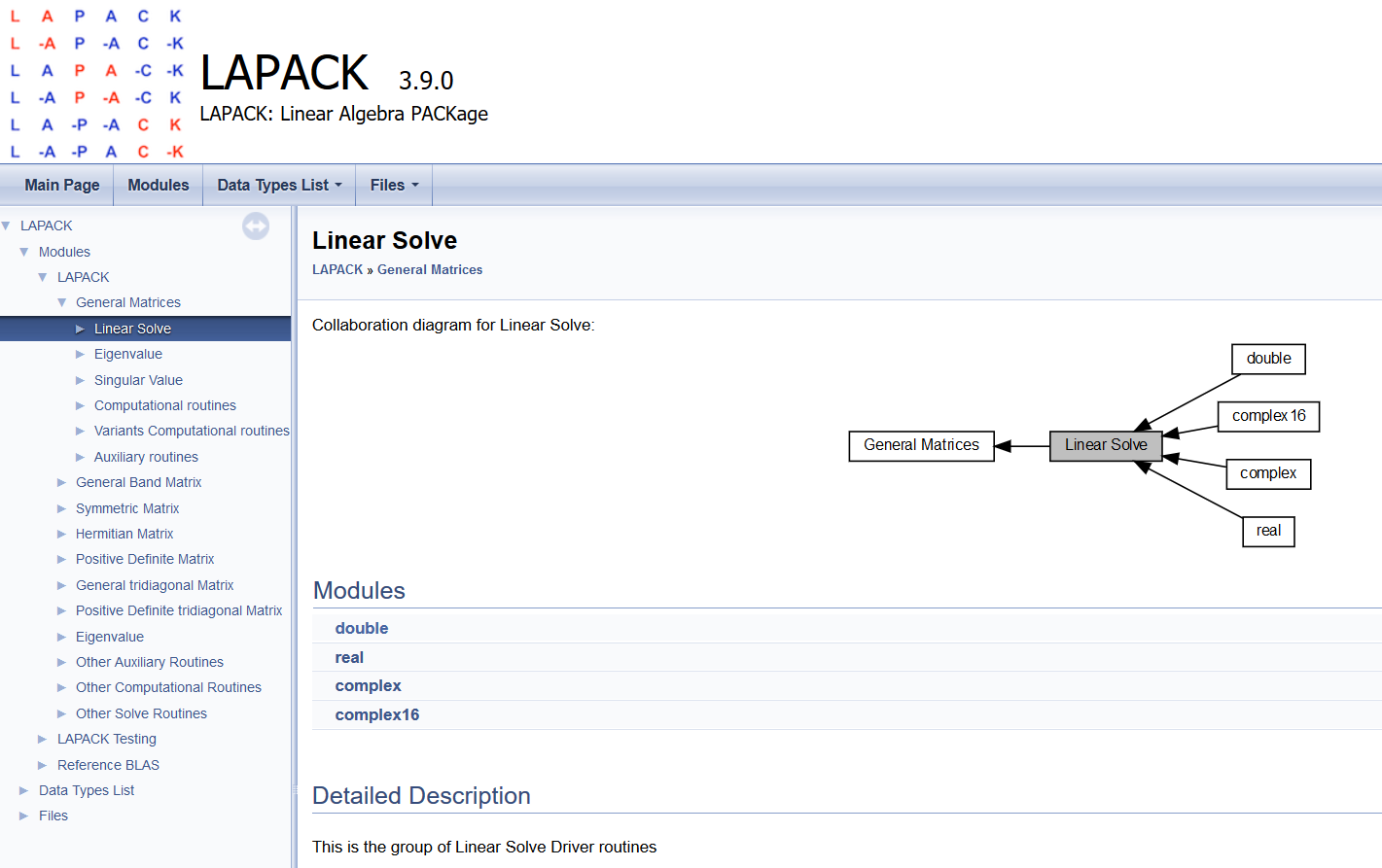
Scipy wrapper API¶
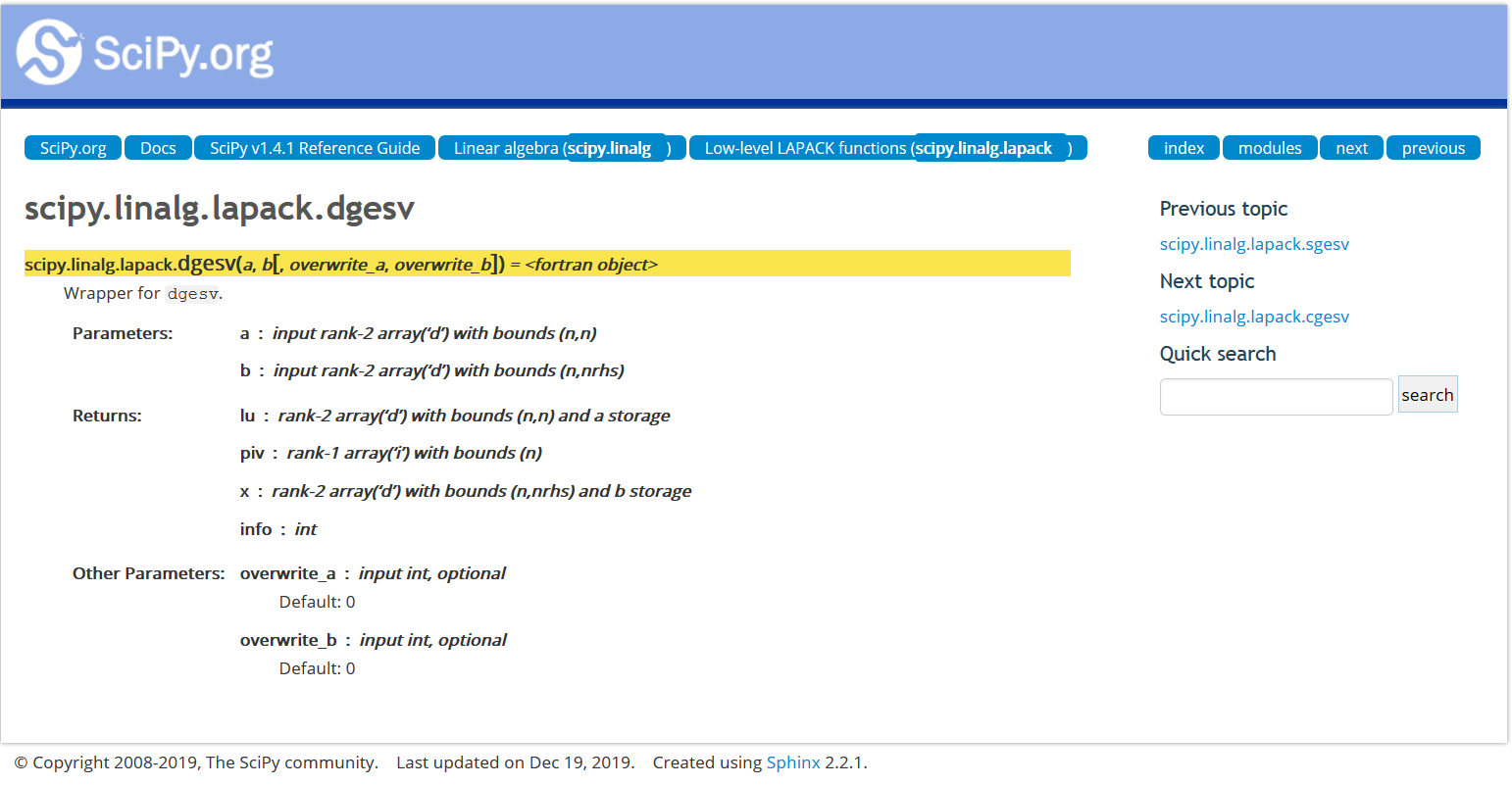
Eigen¶
- https://eigen.tuxfamily.org/dox/
- Written in C++
- Used for linear algebra in Tensorflow (single CPU variant probably)
- Low and high-level (e.g. matrix decompositions) linear algebra functions
- Sparse linear algebra
...name can be annoyingly google-proof however
Take-home message¶
Much (most) code you get for free, or even pay for, that implements mathematical algorithms may be of limited value.
Linear algebra, however, is "solved", in terms of numerical programming.
Your main concern is to find an implementation that fits for your application. E.g. parallel, or sparse, or optimized for a particular CPU.
Lab: Import and directly use packages in Python¶
- BLAS - compute a dot product of two random vectors
- LAPACK - solve a random linear system
II. Overview of leading Python math tools¶
Leading Python tools:¶
Jupyter - "notebooks" for inline code + LaTex math + markup, etc.
NumPy - low-level array & matrix handling and algorithms
SciPy - higher level numerical algorithms (still fairly basic)
Matplotlib - matlab-style plotting & image display
SkLearn - (Scikit-learn) Machine Learning library
NumPy¶
- Numerical algorithm toolbox
- Similar to Matlab but Many key differences, such as zero-based indexing (like C) instead of one-based (like math texts).
- Uses BLAS/LAPACK typically
- NumPy Arrays - special data structure which allows direct and efficient linear algebra manipulations (basically a list with added functionality).
import numpy as np
x = np.array([2,0,1,8])
print(x)
[2 0 1 8]
Fast Numerical Mathematics¶
l = range(1234)
%timeit [i ** 2 for i in l]
389 µs ± 8.15 µs per loop (mean ± std. dev. of 7 runs, 1000 loops each)
import numpy as np
a = np.arange(1234)
%timeit a ** 2
1.41 µs ± 7.46 ns per loop (mean ± std. dev. of 7 runs, 1000000 loops each)
NumPy for Matlab Users¶
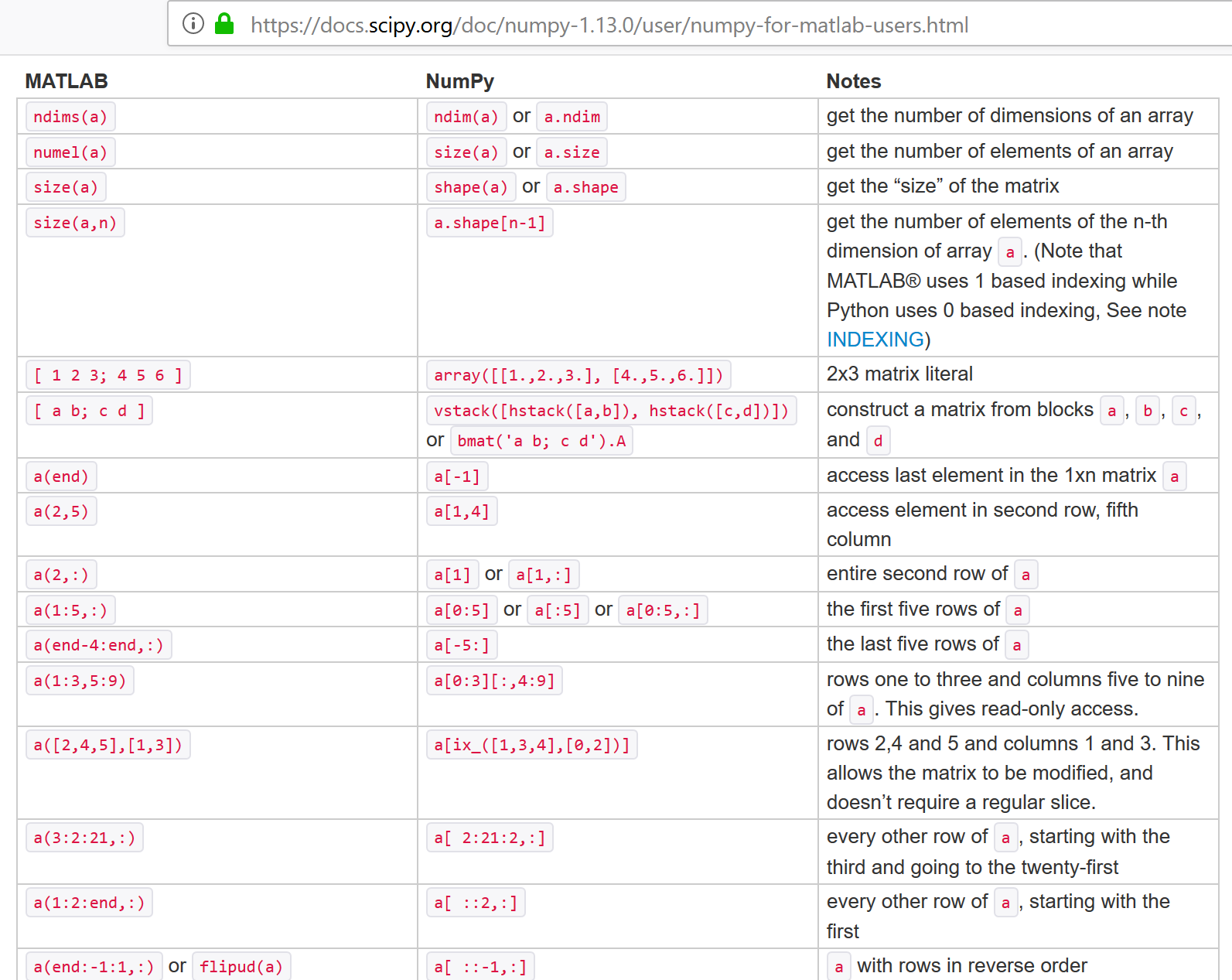
help(np.array)
Help on built-in function array in module numpy.core.multiarray:
array(...)
array(object, dtype=None, copy=True, order='K', subok=False, ndmin=0)
Create an array.
Parameters
----------
object : array_like
An array, any object exposing the array interface, an object whose
__array__ method returns an array, or any (nested) sequence.
dtype : data-type, optional
The desired data-type for the array. If not given, then the type will
be determined as the minimum type required to hold the objects in the
sequence. This argument can only be used to 'upcast' the array. For
downcasting, use the .astype(t) method.
copy : bool, optional
If true (default), then the object is copied. Otherwise, a copy will
only be made if __array__ returns a copy, if obj is a nested sequence,
or if a copy is needed to satisfy any of the other requirements
(`dtype`, `order`, etc.).
order : {'K', 'A', 'C', 'F'}, optional
Specify the memory layout of the array. If object is not an array, the
newly created array will be in C order (row major) unless 'F' is
specified, in which case it will be in Fortran order (column major).
If object is an array the following holds.
===== ========= ===================================================
order no copy copy=True
===== ========= ===================================================
'K' unchanged F & C order preserved, otherwise most similar order
'A' unchanged F order if input is F and not C, otherwise C order
'C' C order C order
'F' F order F order
===== ========= ===================================================
When ``copy=False`` and a copy is made for other reasons, the result is
the same as if ``copy=True``, with some exceptions for `A`, see the
Notes section. The default order is 'K'.
subok : bool, optional
If True, then sub-classes will be passed-through, otherwise
the returned array will be forced to be a base-class array (default).
ndmin : int, optional
Specifies the minimum number of dimensions that the resulting
array should have. Ones will be pre-pended to the shape as
needed to meet this requirement.
Returns
-------
out : ndarray
An array object satisfying the specified requirements.
See Also
--------
empty, empty_like, zeros, zeros_like, ones, ones_like, full, full_like
Notes
-----
When order is 'A' and `object` is an array in neither 'C' nor 'F' order,
and a copy is forced by a change in dtype, then the order of the result is
not necessarily 'C' as expected. This is likely a bug.
Examples
--------
>>> np.array([1, 2, 3])
array([1, 2, 3])
Upcasting:
>>> np.array([1, 2, 3.0])
array([ 1., 2., 3.])
More than one dimension:
>>> np.array([[1, 2], [3, 4]])
array([[1, 2],
[3, 4]])
Minimum dimensions 2:
>>> np.array([1, 2, 3], ndmin=2)
array([[1, 2, 3]])
Type provided:
>>> np.array([1, 2, 3], dtype=complex)
array([ 1.+0.j, 2.+0.j, 3.+0.j])
Data-type consisting of more than one element:
>>> x = np.array([(1,2),(3,4)],dtype=[('a','<i4'),('b','<i4')])
>>> x['a']
array([1, 3])
Creating an array from sub-classes:
>>> np.array(np.mat('1 2; 3 4'))
array([[1, 2],
[3, 4]])
>>> np.array(np.mat('1 2; 3 4'), subok=True)
matrix([[1, 2],
[3, 4]])
Will cover more later, but first a quick intro to some other packages.
SciPy¶
Implements higher-level scientific algorithms using NumPy. Examples:
- Integration (scipy.integrate)
- Optimization (scipy.optimize)
- Interpolation (scipy.interpolate)
- Signal Processing (scipy.signal)
- Linear Algebra (scipy.linalg)
- Statistics (scipy.stats)
- File IO (scipy.io)
Also uses BLAS/LAPACK
import scipy
dir(scipy)
['ALLOW_THREADS', 'AxisError', 'BUFSIZE', 'CLIP', 'ComplexWarning', 'DataSource', 'ERR_CALL', 'ERR_DEFAULT', 'ERR_IGNORE', 'ERR_LOG', 'ERR_PRINT', 'ERR_RAISE', 'ERR_WARN', 'FLOATING_POINT_SUPPORT', 'FPE_DIVIDEBYZERO', 'FPE_INVALID', 'FPE_OVERFLOW', 'FPE_UNDERFLOW', 'False_', 'Inf', 'Infinity', 'LowLevelCallable', 'MAXDIMS', 'MAY_SHARE_BOUNDS', 'MAY_SHARE_EXACT', 'MachAr', 'ModuleDeprecationWarning', 'NAN', 'NINF', 'NZERO', 'NaN', 'PINF', 'PZERO', 'PackageLoader', 'RAISE', 'RankWarning', 'SHIFT_DIVIDEBYZERO', 'SHIFT_INVALID', 'SHIFT_OVERFLOW', 'SHIFT_UNDERFLOW', 'ScalarType', 'TooHardError', 'True_', 'UFUNC_BUFSIZE_DEFAULT', 'UFUNC_PYVALS_NAME', 'VisibleDeprecationWarning', 'WRAP', '__SCIPY_SETUP__', '__all__', '__builtins__', '__cached__', '__config__', '__doc__', '__file__', '__loader__', '__name__', '__numpy_version__', '__package__', '__path__', '__spec__', '__version__', '_distributor_init', '_lib', 'absolute', 'absolute_import', 'add', 'add_docstring', 'add_newdoc', 'add_newdoc_ufunc', 'add_newdocs', 'alen', 'all', 'allclose', 'alltrue', 'amax', 'amin', 'angle', 'any', 'append', 'apply_along_axis', 'apply_over_axes', 'arange', 'arccos', 'arccosh', 'arcsin', 'arcsinh', 'arctan', 'arctan2', 'arctanh', 'argmax', 'argmin', 'argpartition', 'argsort', 'argwhere', 'around', 'array', 'array2string', 'array_equal', 'array_equiv', 'array_repr', 'array_split', 'array_str', 'asanyarray', 'asarray', 'asarray_chkfinite', 'ascontiguousarray', 'asfarray', 'asfortranarray', 'asmatrix', 'asscalar', 'atleast_1d', 'atleast_2d', 'atleast_3d', 'average', 'bartlett', 'base_repr', 'binary_repr', 'bincount', 'bitwise_and', 'bitwise_not', 'bitwise_or', 'bitwise_xor', 'blackman', 'block', 'bmat', 'bool8', 'bool_', 'broadcast', 'broadcast_arrays', 'broadcast_to', 'busday_count', 'busday_offset', 'busdaycalendar', 'byte', 'byte_bounds', 'bytes0', 'bytes_', 'c_', 'can_cast', 'cast', 'cbrt', 'cdouble', 'ceil', 'cfloat', 'char', 'character', 'chararray', 'choose', 'clip', 'clongdouble', 'clongfloat', 'column_stack', 'common_type', 'compare_chararrays', 'complex128', 'complex64', 'complex_', 'complexfloating', 'compress', 'concatenate', 'conj', 'conjugate', 'convolve', 'copy', 'copysign', 'copyto', 'corrcoef', 'correlate', 'cos', 'cosh', 'count_nonzero', 'cov', 'cross', 'csingle', 'ctypeslib', 'cumprod', 'cumproduct', 'cumsum', 'datetime64', 'datetime_as_string', 'datetime_data', 'deg2rad', 'degrees', 'delete', 'deprecate', 'deprecate_with_doc', 'diag', 'diag_indices', 'diag_indices_from', 'diagflat', 'diagonal', 'diff', 'digitize', 'disp', 'divide', 'division', 'divmod', 'dot', 'double', 'dsplit', 'dstack', 'dtype', 'e', 'ediff1d', 'einsum', 'einsum_path', 'emath', 'empty', 'empty_like', 'equal', 'erf', 'errstate', 'euler_gamma', 'exp', 'exp2', 'expand_dims', 'expm1', 'extract', 'eye', 'fabs', 'fastCopyAndTranspose', 'fft', 'fill_diagonal', 'find_common_type', 'finfo', 'fix', 'flatiter', 'flatnonzero', 'flexible', 'flip', 'fliplr', 'flipud', 'float16', 'float32', 'float64', 'float_', 'float_power', 'floating', 'floor', 'floor_divide', 'fmax', 'fmin', 'fmod', 'format_float_positional', 'format_float_scientific', 'format_parser', 'frexp', 'frombuffer', 'fromfile', 'fromfunction', 'fromiter', 'frompyfunc', 'fromregex', 'fromstring', 'full', 'full_like', 'fv', 'generic', 'genfromtxt', 'geomspace', 'get_array_wrap', 'get_include', 'get_printoptions', 'getbufsize', 'geterr', 'geterrcall', 'geterrobj', 'gradient', 'greater', 'greater_equal', 'half', 'hamming', 'hanning', 'heaviside', 'histogram', 'histogram2d', 'histogramdd', 'hsplit', 'hstack', 'hypot', 'i0', 'identity', 'ifft', 'iinfo', 'imag', 'in1d', 'index_exp', 'indices', 'inexact', 'inf', 'info', 'infty', 'inner', 'insert', 'int0', 'int16', 'int32', 'int64', 'int8', 'int_', 'int_asbuffer', 'intc', 'integer', 'interp', 'intersect1d', 'intp', 'invert', 'ipmt', 'irr', 'is_busday', 'isclose', 'iscomplex', 'iscomplexobj', 'isfinite', 'isfortran', 'isin', 'isinf', 'isnan', 'isnat', 'isneginf', 'isposinf', 'isreal', 'isrealobj', 'isscalar', 'issctype', 'issubclass_', 'issubdtype', 'issubsctype', 'iterable', 'ix_', 'kaiser', 'kron', 'ldexp', 'left_shift', 'less', 'less_equal', 'lexsort', 'linspace', 'little_endian', 'load', 'loads', 'loadtxt', 'log', 'log10', 'log1p', 'log2', 'logaddexp', 'logaddexp2', 'logical_and', 'logical_not', 'logical_or', 'logical_xor', 'logn', 'logspace', 'long', 'longcomplex', 'longdouble', 'longfloat', 'longlong', 'lookfor', 'ma', 'mafromtxt', 'mask_indices', 'mat', 'math', 'matmul', 'matrix', 'maximum', 'maximum_sctype', 'may_share_memory', 'mean', 'median', 'memmap', 'meshgrid', 'mgrid', 'min_scalar_type', 'minimum', 'mintypecode', 'mirr', 'mod', 'modf', 'moveaxis', 'msort', 'multiply', 'nan', 'nan_to_num', 'nanargmax', 'nanargmin', 'nancumprod', 'nancumsum', 'nanmax', 'nanmean', 'nanmedian', 'nanmin', 'nanpercentile', 'nanprod', 'nanstd', 'nansum', 'nanvar', 'nbytes', 'ndarray', 'ndenumerate', 'ndfromtxt', 'ndim', 'ndindex', 'nditer', 'negative', 'nested_iters', 'newaxis', 'nextafter', 'nonzero', 'not_equal', 'nper', 'npv', 'number', 'obj2sctype', 'object0', 'object_', 'ogrid', 'ones', 'ones_like', 'outer', 'packbits', 'pad', 'partition', 'percentile', 'pi', 'piecewise', 'pkgload', 'place', 'pmt', 'poly', 'poly1d', 'polyadd', 'polyder', 'polydiv', 'polyfit', 'polyint', 'polymul', 'polysub', 'polyval', 'positive', 'power', 'ppmt', 'print_function', 'prod', 'product', 'promote_types', 'ptp', 'put', 'putmask', 'pv', 'r_', 'rad2deg', 'radians', 'rand', 'randn', 'random', 'rank', 'rate', 'ravel', 'ravel_multi_index', 'real', 'real_if_close', 'rec', 'recarray', 'recfromcsv', 'recfromtxt', 'reciprocal', 'record', 'remainder', 'repeat', 'require', 'reshape', 'resize', 'result_type', 'right_shift', 'rint', 'roll', 'rollaxis', 'roots', 'rot90', 'round_', 'row_stack', 's_', 'safe_eval', 'save', 'savetxt', 'savez', 'savez_compressed', 'sctype2char', 'sctypeDict', 'sctypeNA', 'sctypes', 'searchsorted', 'select', 'set_numeric_ops', 'set_printoptions', 'set_string_function', 'setbufsize', 'setdiff1d', 'seterr', 'seterrcall', 'seterrobj', 'setxor1d', 'shape', 'shares_memory', 'short', 'show_config', 'show_numpy_config', 'sign', 'signbit', 'signedinteger', 'sin', 'sinc', 'single', 'singlecomplex', 'sinh', 'size', 'sometrue', 'sort', 'sort_complex', 'source', 'spacing', 'split', 'sqrt', 'square', 'squeeze', 'stack', 'std', 'str0', 'str_', 'string_', 'subtract', 'sum', 'swapaxes', 'take', 'tan', 'tanh', 'tensordot', 'test', 'tile', 'timedelta64', 'trace', 'tracemalloc_domain', 'transpose', 'trapz', 'tri', 'tril', 'tril_indices', 'tril_indices_from', 'trim_zeros', 'triu', 'triu_indices', 'triu_indices_from', 'true_divide', 'trunc', 'typeDict', 'typeNA', 'typecodes', 'typename', 'ubyte', 'ufunc', 'uint', 'uint0', 'uint16', 'uint32', 'uint64', 'uint8', 'uintc', 'uintp', 'ulonglong', 'unicode', 'unicode_', 'union1d', 'unique', 'unpackbits', 'unravel_index', 'unsignedinteger', 'unwrap', 'ushort', 'vander', 'var', 'vdot', 'vectorize', 'version', 'void', 'void0', 'vsplit', 'vstack', 'where', 'who', 'zeros', 'zeros_like']
Matplotlib¶
Tutorial from: https://github.com/amueller/scipy-2017-sklearn/blob/master/notebooks/02.Scientific_Computing_Tools_in_Python.ipynb
Another important part of machine learning is the visualization of data. The most common
tool for this in Python is matplotlib. It is an extremely flexible package, and
we will go over some basics here.
Jupyter has built-in "magic functions", the "matoplotlib inline" mode, which will draw the plots directly inside the notebook. Should be on by default.
%matplotlib inline
import matplotlib.pyplot as plt
# Plotting a line
x = np.linspace(0, 10, 100)
plt.plot(x, np.sin(x));
# Scatter-plot points
x = np.random.normal(size=500)
y = np.random.normal(size=500)
plt.scatter(x, y);
# Showing images using imshow
# - note that origin is at the top-left by default!
#x = np.linspace(1, 12, 100)
#y = x[:, np.newaxis]
#im = y * np.sin(x) * np.cos(y)
im = np.array([[1, 2, 3],[4,5,6],[6,7,8]])
import matplotlib.pyplot as plt
plt.imshow(im);
plt.colorbar();
plt.xlabel('x')
plt.ylabel('y')
plt.show();
# Contour plots
# - note that origin here is at the bottom-left by default!
plt.contour(im);
# 3D plotting
from mpl_toolkits.mplot3d import Axes3D
ax = plt.axes(projection='3d')
xgrid, ygrid = np.meshgrid(x, y.ravel())
ax.plot_surface(xgrid, ygrid, im, cmap=plt.cm.viridis, cstride=2, rstride=2, linewidth=0);
There are many more plot types available. See matplotlib gallery.
Test these examples: copy the Source Code
link, and put it in a notebook using the %load magic.
For example:
# %load http://matplotlib.org/mpl_examples/pylab_examples/ellipse_collection.py
SkLearn¶
- Many Machine Learning functions.
- Couple dozen core developers + hundreds of other contributors.
- 2011 tutorial has over 10,000 citations.
"Scikit-learn: Machine Learning in Python", Fabian Pedregosa, Gaël Varoquaux, Alexandre Gramfort, Vincent Michel, Bertrand Thirion, Olivier Grisel, Mathieu Blondel, Peter Prettenhofer, Ron Weiss, Vincent Dubourg, Jake Vanderplas, Alexandre Passos, David Cournapeau, Matthieu Brucher, Matthieu Perrot, Édouard Duchesnay; 12(Oct):2825−2830, 2011
Considered leading Machine Learning toolbox. Was getting discarded as field switched from sinlge CPU to multicore, but appears to be making comeback with parallel upgrades
import sklearn as sk
dir(sk)
['_ASSUME_FINITE', '__SKLEARN_SETUP__', '__all__', '__builtins__', '__cached__', '__check_build', '__doc__', '__file__', '__loader__', '__name__', '__package__', '__path__', '__spec__', '__version__', '_contextmanager', '_isotonic', 'base', 'calibration', 'clone', 'cluster', 'config_context', 'covariance', 'cross_decomposition', 'cross_validation', 'datasets', 'decomposition', 'discriminant_analysis', 'dummy', 'ensemble', 'exceptions', 'externals', 'feature_extraction', 'feature_selection', 'gaussian_process', 'get_config', 'grid_search', 'isotonic', 'kernel_approximation', 'kernel_ridge', 'learning_curve', 'linear_model', 'logger', 'logging', 'manifold', 'metrics', 'mixture', 'model_selection', 'multiclass', 'multioutput', 'naive_bayes', 'neighbors', 'neural_network', 'os', 'pipeline', 'preprocessing', 'random_projection', 're', 'semi_supervised', 'set_config', 'setup_module', 'svm', 'sys', 'tree', 'utils', 'warnings']
Machine Learning Framework using Sklearn¶
Given training data $(\mathbf x_{(i)},y_i)$ for $i=1,...,m$ --> lists of samples $\verb|X|$ and labels $\verb|y|$
Choose a model $f(\cdot)$ where we want to make $f(\mathbf x_{(i)})\approx y_i$ (for all $i$) --> choose sklearn estimator to use
Define a loss function $L(f(\mathbf x), y)$ to minimize by changing $f(\cdot)$ ...by adjusting the weights --> default choises for estimators, sometimes multiple options
The Sklearn API¶
sklearn has an Object Oriented interface
Most models/transforms/objects in sklearn are Estimator objects
class Estimator(object):
def fit(self, X, y=None):
"""Fit model to data X (and y)"""
self.some_attribute = self.some_fitting_method(X, y)
return self
def predict(self, X_test):
"""Make prediction based on passed features"""
pred = self.make_prediction(X_test)
return pred
model = Estimator()
The Estimator class defines a fit() method as well as a predict() method. For an instance of an Estimator stored in a variable model:
model.fit: fits the model with the passed in training data. For supervised models, it also accepts a second argumentythat corresponds to the labels (model.fit(X, y). For unsupervised models, there are no labels so you only need to pass in the feature matrix (model.fit(X))Since the interface is very OO, the instance itself stores the results of the
fitinternally. And as such you must alwaysfit()before youpredict()on the same object.model.predict: predicts new labels for any new datapoints passed in (model.predict(X_test)) and returns an array equal in length to the number of rows of what is passed in containing the predicted labels.
Types of subclass of estimator:¶
- Supervised
- Unsupervised
- Feature Processing
Supervised¶
Supervised estimators in addition to the above methods typically also have:
model.predict_proba: For classifiers that have a notion of probability (or some measure of confidence in a prediction) this method returns those "probabilities". The label with the highest probability is what is returned by themodel.predict()` mehod from above.model.score: For both classification and regression models, this method returns some measure of validation of the model (which is configurable). For example, in regression the default is typically R^2 and classification it is accuracy.
Unsupervised - Transformer interface¶
Some estimators in the library implement this.
Unsupervised in this case refers to any method that does not need labels, including unsupervised classifiers, preprocessing (like tf-idf), dimensionality reduction, etc.
The transformer interface usually defines two additional methods:
model.transform: Given an unsupervised model, transform the input into a new basis (or feature space). This accepts on argument (usually a feature matrix) and returns a matrix of the input transformed. Note: You need tofit()the model before you transform it.model.fit_transform: For some models you may not need tofit()andtransform()separately. In these cases it is more convenient to do both at the same time.
X = [[0], [1], [2], [3]]
y = [0, 0, 1, 1]
from sklearn.neighbors import KNeighborsClassifier
neigh = KNeighborsClassifier(n_neighbors=3)
neigh.fit(X, y)
print(neigh.predict([[1.1]]))
print(neigh.predict_proba([[0.9]]))
[0] [[0.66666667 0.33333333]]
III. Structured and Sparse Matrices¶
Motivation¶
If you know your matric has particular structure, you can take advantage of this to greatly reduce computations and/or storage
Examples:
- Diagonal matrix
- Banded matrix
- Toeplitz (or circulant) matrix --> convolutional neural nets
- Sparse matrix
- Kronecker product
Structured matrix examples¶
How would you efficiently store these?

How might you efficiently multiple by a vector?
Examples¶
- Sparse structured matrices for which we don't have a specialized implementation
- Matrices describing sparse networks
scipy.sparse¶
SciPy 2-D sparse matrix package for numeric data.
https://docs.scipy.org/doc/scipy/reference/generated/scipy.sparse.csr_matrix.html
Sparse matrix types:
- bsr_matrix - Block Sparse Row matrix
- coo_matrix = A sparse matrix in COOrdinate format.
- csc_matrix - Compressed Sparse Column matrix
- csr_matrix - Compressed Sparse Row matrix
- dia_matrix - Sparse matrix with DIAgonal storage
- dok_matrix - Dictionary Of Keys based sparse matrix.
- lil_matrix - Row-based linked list sparse matrix
Different formats will be faster or slower for different tasks. Read API.
Compressed Sparse Row (CSR) Format¶
Stores list of values, list of column indices, and value-list indices for values that start each row
Also known as Compressed Row Storage CRS
from scipy.sparse import csr_matrix
indptr = np.array([0, 2, 3, 6])
indices = np.array([0, 2, 2, 0, 1, 2])
data = np.array([1, 2, 3, 4, 5, 6])
csr_matrix((data, indices, indptr), shape=(3, 3)).toarray()
array([[1, 0, 2],
[0, 0, 3],
[4, 5, 6]])
# can also define with simple indices
row = np.array([0, 0, 1, 2, 2, 2])
col = np.array([0, 2, 2, 0, 1, 2])
data = np.array([1, 2, 3, 4, 5, 6])
csr_matrix((data, (row, col)), shape=(3, 3)).toarray()
array([[1, 0, 2],
[0, 0, 3],
[4, 5, 6]], dtype=int32)
Sparse solvers¶
- High-level algorithms that convert operations into BLAS calculations
- More limited implementations - sometimes no one yet happened to adapt a sparse solver to your particular problem
Exercise¶
Create two random matrices then make them sparse by zeroing out most elements (e.g. $\verb|A[A<0.99]=0|$)
Compare speed of matrix multiplication for different sparse formats
Recap¶
- Mathematical software is implemented hierarchically
- BLAS = Lowest level(s), implemented on particular hardware, e.g. for SIMD processors
- Higher level functions (solvers and decompositions) calls BLAS to implement internal computations
- Structured and Sparse matrix libraries also call BLAS typically (convert large sparse problems to small dense problems)
- Modern alternatives also can utilize BLAS/LAPACK backend and also suppot hardware-specific replacements (e.g. CUDA)
IV. NumPy Basics¶
NumPy Arrays for Vectors¶
x = np.array([2,0,1,8])
x
array([2, 0, 1, 8])
NumPy arrays are zero-based like Python lists, but linear algebra writings are typically one-based.
print(x[0],x[3])
2 8
What is a "Tensor"? (Computer Science version)¶
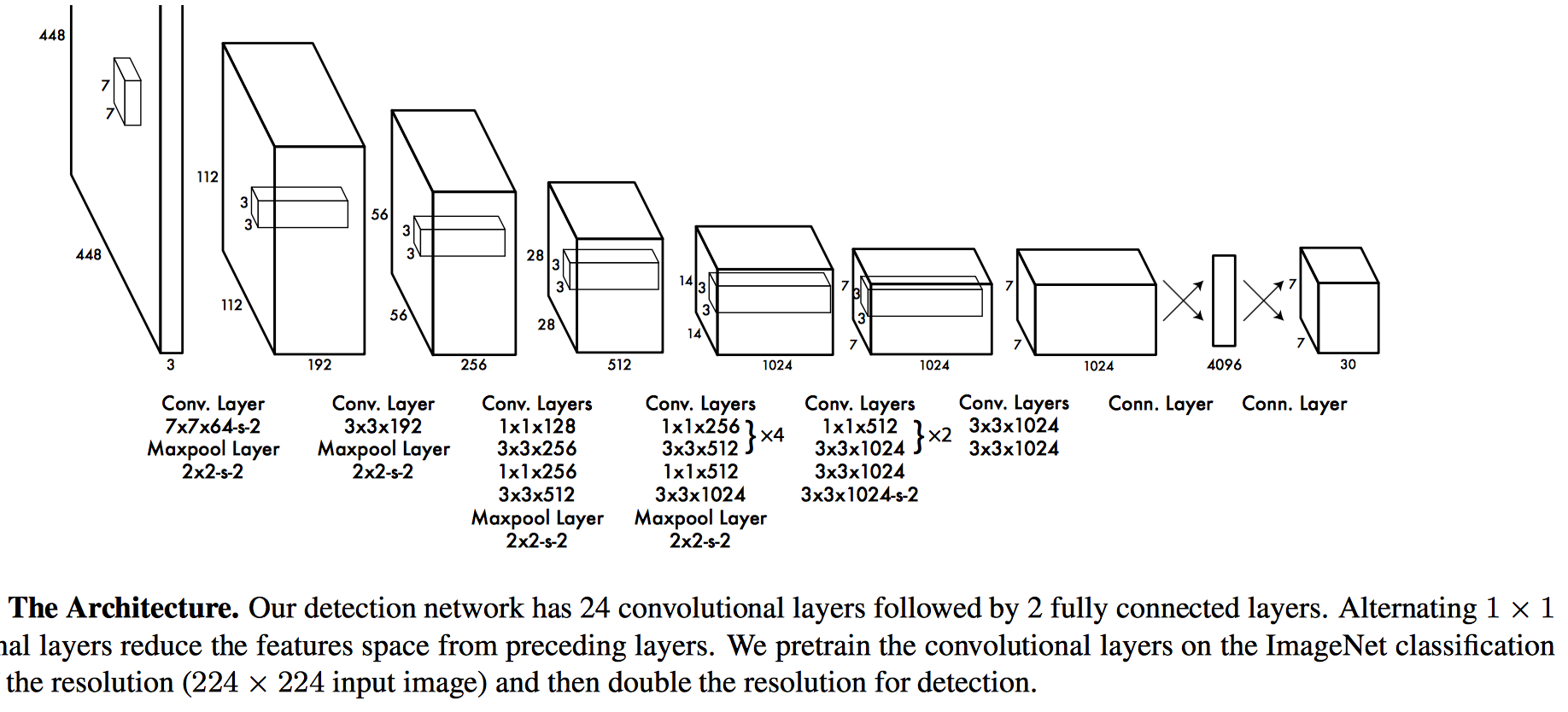

Behind the scenes it's just another list of numbers with some extra info regarding dimensions
Ndarray¶
https://docs.scipy.org/doc/numpy-1.13.0/reference/arrays.ndarray.html
A $d$-dimensional data structure, containing $n_1\times n_2 \times ... \times n_d$ numbers
np.random.rand(3)
array([0.57100166, 0.07586713, 0.90062779])
np.random.rand(3,2) # note not a tuple (unlike many other numpy functions)
array([[0.49662458, 0.23158612],
[0.94269648, 0.84962285],
[0.68847847, 0.73942405]])
np.random.rand(3,2,4)
array([[[0.09840227, 0.26310991, 0.58589047, 0.33265107],
[0.89329091, 0.99117784, 0.49475921, 0.08047502]],
[[0.09570909, 0.28074458, 0.15119848, 0.99682941],
[0.46506878, 0.0333583 , 0.08488412, 0.26735574]],
[[0.65350538, 0.43238726, 0.79292367, 0.58734204],
[0.3019322 , 0.21985196, 0.41835067, 0.44614492]]])
- A list of length 3,
- each of those 3 elements is a list of length 2
- each of those 2 elements is a list of length 4
T = np.random.rand(3,2,4,3,2)
T.shape
(3, 2, 4, 3, 2)
T.ndim
5
$\verb|numpy.linalg|$¶
- Use BLAS and LAPACK to provide efficient low level implementations of standard linear algebra algorithms.
- Those libraries may be provided by NumPy itself using C versions of a subset of their reference implementations
- When possible, highly optimized libraries that take advantage of specialized processor functionality are preferred.
- Examples of such libraries are OpenBLAS, MKL (TM), and ATLAS.
https://docs.scipy.org/doc/numpy/reference/routines.linalg.html
Matrix and vector products¶
- dot(a, b[, out]) Dot product of two arrays.
- linalg.multi_dot(arrays) Compute the dot product of two or more arrays in a single function call, while automatically selecting the fastest evaluation order.
- vdot(a, b) Return the dot product of two vectors.
- inner(a, b) Inner product of two arrays.
- outer(a, b[, out]) Compute the outer product of two vectors.
- matmul(x1, x2, /[, out, casting, order, …]) Matrix product of two arrays.
- tensordot(a, b[, axes]) Compute tensor dot product along specified axes.
- einsum(subscripts, *operands[, out, dtype, …]) Evaluates the Einstein summation convention on the operands.
- einsum_path(subscripts, *operands[, optimize]) Evaluates the lowest cost contraction order for an einsum expression by considering the creation of intermediate arrays.
- linalg.matrix_power(a, n) Raise a square matrix to the (integer) power n.
- kron(a, b) Kronecker product of two arrays.
Decompositions¶
- linalg.cholesky(a) Cholesky decomposition.
- linalg.qr(a[, mode]) Compute the qr factorization of a matrix.
- linalg.svd(a[, full_matrices, compute_uv, …]) Singular Value Decomposition.
Matrix eigenvalues¶¶
- linalg.eig(a) Compute the eigenvalues and right eigenvectors of a square array.
- linalg.eigh(a[, UPLO]) Return the eigenvalues and eigenvectors of a complex Hermitian (conjugate symmetric) or a real symmetric matrix.
- linalg.eigvals(a) Compute the eigenvalues of a general matrix.
- linalg.eigvalsh(a[, UPLO]) Compute the eigenvalues of a complex Hermitian or real symmetric matrix.
Norms and other numbers¶
- linalg.norm(x[, ord, axis, keepdims]) Matrix or vector norm.
- linalg.cond(x[, p]) Compute the condition number of a matrix.
- linalg.det(a) Compute the determinant of an array.
- linalg.matrix_rank(M[, tol, hermitian]) Return matrix rank of array using SVD method
- linalg.slogdet(a) Compute the sign and (natural) logarithm of the determinant of an array.
- trace(a[, offset, axis1, axis2, dtype, out]) Return the sum along diagonals of the array.
Solving equations and inverting matrices¶
- linalg.solve(a, b) Solve a linear matrix equation, or system of linear scalar equations.
- linalg.tensorsolve(a, b[, axes]) Solve the tensor equation a x = b for x.
- linalg.lstsq(a, b[, rcond]) Return the least-squares solution to a linear matrix equation.
- linalg.inv(a) Compute the (multiplicative) inverse of a matrix.
- linalg.pinv(a[, rcond, hermitian]) Compute the (Moore-Penrose) pseudo-inverse of a matrix.
- linalg.tensorinv(a[, ind]) Compute the ‘inverse’ of an N-dimensional array.
"Vectorization"¶
Converting loops into linear algebra functions where loop is performed at lower level
E.g. instead of looping over a list to compute squares of elements, make into array and "square" array
v = [1,2,3,4,5]
v2 = []
for v_k in v:
v2.append(v_k**2)
v2
[1, 4, 9, 16, 25]
np.array(v)**2
array([ 1, 4, 9, 16, 25], dtype=int32)
Dot Product using NumPy¶
w = np.array([1, 2])
v = np.array([3, 4])
np.dot(w,v)
11
w = np.array([0, 1])
v = np.array([1, 0])
np.dot(w,v)
0
w = np.array([1, 2])
v = np.array([-2, 1])
np.dot(w,v)
0
Matrix-vector multiplication using NumPy - CAUTION¶
NumPy implements a broadcast version of multiplication when using the "*" operator.
It guesses what you mean rather by seeking ways the shapes match, rather than giving error.
w = np.array([[2,-6],[-1, 4]])
v = np.array([12,46])
w*v
array([[ 24, -276],
[ -12, 184]])
v+w
array([[14, 40],
[11, 50]])
"Broadcasting"¶
Multiply matrices or vectrors in an element-wise manner by repeating "singular" dimensions.
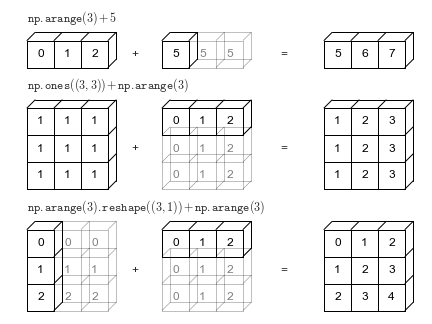
Matlab and Numpy automatically do this now.
Also (and previously) in matlab, e.g.: "brcm(@plus, A, x). numpy.broadcast
For true "Matrix-vector" multiplication, use matmul or "@"¶
a special case of matrix-matrix multiplication
w = np.array([[ 2, -6],
[-1, 4]])
v = np.array([[+1],
[-1]])
np.matmul(w,v)
array([[ 8],
[-5]])
w@v
array([ 8, -5])
Matrix-matrix multiplication¶
A = np.array([[1,2],[3, 4]])
B = np.array([[1,0],[1, 1]])
A*B
array([[1, 0],
[3, 4]])
What is this doing?¶
np.matmul(A,B)
array([[3, 2],
[7, 4]])
np.dot(A,B)
array([[3, 2],
[7, 4]])
A@B
array([[3, 2],
[7, 4]])
Reshaping vectors into matrices and whatever else you want¶
x = np.array([1,2,3,4])
x = x.reshape(4,1)
print(x.reshape(4,1))
[[1] [2] [3] [4]]
print(x.reshape(2,2))
[[1 2] [3 4]]
print(x)
x_T = x.transpose()
print (x_T)
print (x_T.shape)
print(x.T)
[[1] [2] [3] [4]] [[1 2 3 4]] (1, 4) [[1 2 3 4]]
Famous matrices in NumPy¶
np.eye(3)
array([[1., 0., 0.],
[0., 1., 0.],
[0., 0., 1.]])
D = np.diag([1,2,3])
print(D)
[[1 0 0] [0 2 0] [0 0 3]]
np.diag(D)
array([1, 2, 3])
NumPy Inverse¶
A = np.array([[1,2],[4,5]])
iA = np.linalg.inv(A)
print(iA)
print(np.matmul(A,iA))
[[-1.66666667 0.66666667] [ 1.33333333 -0.33333333]] [[1. 0.] [0. 1.]]
2.*8.
16.0
A = np.random.randint(0, 10, size=(3, 3))
A_inv = np.linalg.inv(A)
print(A)
print(A_inv)
print(np.matmul(A,A_inv))
[[2 6 4] [5 7 1] [5 3 2]] [[-1.25000000e-01 -1.11022302e-17 2.50000000e-01] [ 5.68181818e-02 1.81818182e-01 -2.04545455e-01] [ 2.27272727e-01 -2.72727273e-01 1.81818182e-01]] [[ 1.00000000e+00 0.00000000e+00 -1.11022302e-16] [-2.77555756e-17 1.00000000e+00 -8.32667268e-17] [-5.55111512e-17 0.00000000e+00 1.00000000e+00]]
Note that "hard zeros" rarely exist in real world due to limited numerical precision.
Eigenvalue Decomposition in NumPy¶
A = np.asarray([[.3,.1,.6],[.1,.3,.6],[.15,.15,.70]])
np.linalg.eig(A)
(array([1. , 0.2, 0.1]),
array([[-5.77350269e-01, -7.07106781e-01, -6.66666667e-01],
[-5.77350269e-01, 7.07106781e-01, -6.66666667e-01],
[-5.77350269e-01, -2.74027523e-16, 3.33333333e-01]]))
Note there's only one return
(u,s) = np.linalg.eig(A)
print(u)
print(s)
[1. 0.2 0.1] [[-5.77350269e-01 -7.07106781e-01 -6.66666667e-01] [-5.77350269e-01 7.07106781e-01 -6.66666667e-01] [-5.77350269e-01 -2.74027523e-16 3.33333333e-01]]
help(np.linalg.eig)
Help on function eig in module numpy.linalg.linalg:
eig(a)
Compute the eigenvalues and right eigenvectors of a square array.
Parameters
----------
a : (..., M, M) array
Matrices for which the eigenvalues and right eigenvectors will
be computed
Returns
-------
w : (..., M) array
The eigenvalues, each repeated according to its multiplicity.
The eigenvalues are not necessarily ordered. The resulting
array will be of complex type, unless the imaginary part is
zero in which case it will be cast to a real type. When `a`
is real the resulting eigenvalues will be real (0 imaginary
part) or occur in conjugate pairs
v : (..., M, M) array
The normalized (unit "length") eigenvectors, such that the
column ``v[:,i]`` is the eigenvector corresponding to the
eigenvalue ``w[i]``.
Raises
------
LinAlgError
If the eigenvalue computation does not converge.
See Also
--------
eigvals : eigenvalues of a non-symmetric array.
eigh : eigenvalues and eigenvectors of a symmetric or Hermitian
(conjugate symmetric) array.
eigvalsh : eigenvalues of a symmetric or Hermitian (conjugate symmetric)
array.
Notes
-----
.. versionadded:: 1.8.0
Broadcasting rules apply, see the `numpy.linalg` documentation for
details.
This is implemented using the _geev LAPACK routines which compute
the eigenvalues and eigenvectors of general square arrays.
The number `w` is an eigenvalue of `a` if there exists a vector
`v` such that ``dot(a,v) = w * v``. Thus, the arrays `a`, `w`, and
`v` satisfy the equations ``dot(a[:,:], v[:,i]) = w[i] * v[:,i]``
for :math:`i \in \{0,...,M-1\}`.
The array `v` of eigenvectors may not be of maximum rank, that is, some
of the columns may be linearly dependent, although round-off error may
obscure that fact. If the eigenvalues are all different, then theoretically
the eigenvectors are linearly independent. Likewise, the (complex-valued)
matrix of eigenvectors `v` is unitary if the matrix `a` is normal, i.e.,
if ``dot(a, a.H) = dot(a.H, a)``, where `a.H` denotes the conjugate
transpose of `a`.
Finally, it is emphasized that `v` consists of the *right* (as in
right-hand side) eigenvectors of `a`. A vector `y` satisfying
``dot(y.T, a) = z * y.T`` for some number `z` is called a *left*
eigenvector of `a`, and, in general, the left and right eigenvectors
of a matrix are not necessarily the (perhaps conjugate) transposes
of each other.
References
----------
G. Strang, *Linear Algebra and Its Applications*, 2nd Ed., Orlando, FL,
Academic Press, Inc., 1980, Various pp.
Examples
--------
>>> from numpy import linalg as LA
(Almost) trivial example with real e-values and e-vectors.
>>> w, v = LA.eig(np.diag((1, 2, 3)))
>>> w; v
array([ 1., 2., 3.])
array([[ 1., 0., 0.],
[ 0., 1., 0.],
[ 0., 0., 1.]])
Real matrix possessing complex e-values and e-vectors; note that the
e-values are complex conjugates of each other.
>>> w, v = LA.eig(np.array([[1, -1], [1, 1]]))
>>> w; v
array([ 1. + 1.j, 1. - 1.j])
array([[ 0.70710678+0.j , 0.70710678+0.j ],
[ 0.00000000-0.70710678j, 0.00000000+0.70710678j]])
Complex-valued matrix with real e-values (but complex-valued e-vectors);
note that a.conj().T = a, i.e., a is Hermitian.
>>> a = np.array([[1, 1j], [-1j, 1]])
>>> w, v = LA.eig(a)
>>> w; v
array([ 2.00000000e+00+0.j, 5.98651912e-36+0.j]) # i.e., {2, 0}
array([[ 0.00000000+0.70710678j, 0.70710678+0.j ],
[ 0.70710678+0.j , 0.00000000+0.70710678j]])
Be careful about round-off error!
>>> a = np.array([[1 + 1e-9, 0], [0, 1 - 1e-9]])
>>> # Theor. e-values are 1 +/- 1e-9
>>> w, v = LA.eig(a)
>>> w; v
array([ 1., 1.])
array([[ 1., 0.],
[ 0., 1.]])
SVD in NumPy¶
np.linalg.svd(A)
(array([[-0.32453643, -0.9458732 ],
[-0.9458732 , 0.32453643]]),
array([6.76782894, 0.44327362]),
array([[-0.60699365, -0.79470668],
[ 0.79470668, -0.60699365]]))
Note outputs are U, S, and V-transposed
help(np.linalg.svd)
Help on function svd in module numpy.linalg.linalg:
svd(a, full_matrices=True, compute_uv=True)
Singular Value Decomposition.
When `a` is a 2D array, it is factorized as ``u @ np.diag(s) @ vh
= (u * s) @ vh``, where `u` and `vh` are 2D unitary arrays and `s` is a 1D
array of `a`'s singular values. When `a` is higher-dimensional, SVD is
applied in stacked mode as explained below.
Parameters
----------
a : (..., M, N) array_like
A real or complex array with ``a.ndim >= 2``.
full_matrices : bool, optional
If True (default), `u` and `vh` have the shapes ``(..., M, M)`` and
``(..., N, N)``, respectively. Otherwise, the shapes are
``(..., M, K)`` and ``(..., K, N)``, respectively, where
``K = min(M, N)``.
compute_uv : bool, optional
Whether or not to compute `u` and `vh` in addition to `s`. True
by default.
Returns
-------
u : { (..., M, M), (..., M, K) } array
Unitary array(s). The first ``a.ndim - 2`` dimensions have the same
size as those of the input `a`. The size of the last two dimensions
depends on the value of `full_matrices`. Only returned when
`compute_uv` is True.
s : (..., K) array
Vector(s) with the singular values, within each vector sorted in
descending order. The first ``a.ndim - 2`` dimensions have the same
size as those of the input `a`.
vh : { (..., N, N), (..., K, N) } array
Unitary array(s). The first ``a.ndim - 2`` dimensions have the same
size as those of the input `a`. The size of the last two dimensions
depends on the value of `full_matrices`. Only returned when
`compute_uv` is True.
Raises
------
LinAlgError
If SVD computation does not converge.
Notes
-----
.. versionchanged:: 1.8.0
Broadcasting rules apply, see the `numpy.linalg` documentation for
details.
The decomposition is performed using LAPACK routine ``_gesdd``.
SVD is usually described for the factorization of a 2D matrix :math:`A`.
The higher-dimensional case will be discussed below. In the 2D case, SVD is
written as :math:`A = U S V^H`, where :math:`A = a`, :math:`U= u`,
:math:`S= \mathtt{np.diag}(s)` and :math:`V^H = vh`. The 1D array `s`
contains the singular values of `a` and `u` and `vh` are unitary. The rows
of `vh` are the eigenvectors of :math:`A^H A` and the columns of `u` are
the eigenvectors of :math:`A A^H`. In both cases the corresponding
(possibly non-zero) eigenvalues are given by ``s**2``.
If `a` has more than two dimensions, then broadcasting rules apply, as
explained in :ref:`routines.linalg-broadcasting`. This means that SVD is
working in "stacked" mode: it iterates over all indices of the first
``a.ndim - 2`` dimensions and for each combination SVD is applied to the
last two indices. The matrix `a` can be reconstructed from the
decomposition with either ``(u * s[..., None, :]) @ vh`` or
``u @ (s[..., None] * vh)``. (The ``@`` operator can be replaced by the
function ``np.matmul`` for python versions below 3.5.)
If `a` is a ``matrix`` object (as opposed to an ``ndarray``), then so are
all the return values.
Examples
--------
>>> a = np.random.randn(9, 6) + 1j*np.random.randn(9, 6)
>>> b = np.random.randn(2, 7, 8, 3) + 1j*np.random.randn(2, 7, 8, 3)
Reconstruction based on full SVD, 2D case:
>>> u, s, vh = np.linalg.svd(a, full_matrices=True)
>>> u.shape, s.shape, vh.shape
((9, 9), (6,), (6, 6))
>>> np.allclose(a, np.dot(u[:, :6] * s, vh))
True
>>> smat = np.zeros((9, 6), dtype=complex)
>>> smat[:6, :6] = np.diag(s)
>>> np.allclose(a, np.dot(u, np.dot(smat, vh)))
True
Reconstruction based on reduced SVD, 2D case:
>>> u, s, vh = np.linalg.svd(a, full_matrices=False)
>>> u.shape, s.shape, vh.shape
((9, 6), (6,), (6, 6))
>>> np.allclose(a, np.dot(u * s, vh))
True
>>> smat = np.diag(s)
>>> np.allclose(a, np.dot(u, np.dot(smat, vh)))
True
Reconstruction based on full SVD, 4D case:
>>> u, s, vh = np.linalg.svd(b, full_matrices=True)
>>> u.shape, s.shape, vh.shape
((2, 7, 8, 8), (2, 7, 3), (2, 7, 3, 3))
>>> np.allclose(b, np.matmul(u[..., :3] * s[..., None, :], vh))
True
>>> np.allclose(b, np.matmul(u[..., :3], s[..., None] * vh))
True
Reconstruction based on reduced SVD, 4D case:
>>> u, s, vh = np.linalg.svd(b, full_matrices=False)
>>> u.shape, s.shape, vh.shape
((2, 7, 8, 3), (2, 7, 3), (2, 7, 3, 3))
>>> np.allclose(b, np.matmul(u * s[..., None, :], vh))
True
>>> np.allclose(b, np.matmul(u, s[..., None] * vh))
True
QR in NumPy¶
Q,R = np.linalg.qr(A)
print(Q)
#print(R)
[[-0.85714286 0.39439831 -0.33129458] [-0.28571429 -0.89922814 -0.33129458] [-0.42857143 -0.18931119 0.88345221]]
np.dot(Q[:,2],Q[:,2])
1.0
Q.T@Q
array([[ 1.00000000e+00, 3.25682983e-17, -6.47535828e-17],
[ 3.25682983e-17, 1.00000000e+00, 1.03449228e-16],
[-6.47535828e-17, 1.03449228e-16, 1.00000000e+00]])
help(np.linalg.qr)
Help on function qr in module numpy.linalg.linalg:
qr(a, mode='reduced')
Compute the qr factorization of a matrix.
Factor the matrix `a` as *qr*, where `q` is orthonormal and `r` is
upper-triangular.
Parameters
----------
a : array_like, shape (M, N)
Matrix to be factored.
mode : {'reduced', 'complete', 'r', 'raw', 'full', 'economic'}, optional
If K = min(M, N), then
* 'reduced' : returns q, r with dimensions (M, K), (K, N) (default)
* 'complete' : returns q, r with dimensions (M, M), (M, N)
* 'r' : returns r only with dimensions (K, N)
* 'raw' : returns h, tau with dimensions (N, M), (K,)
* 'full' : alias of 'reduced', deprecated
* 'economic' : returns h from 'raw', deprecated.
The options 'reduced', 'complete, and 'raw' are new in numpy 1.8,
see the notes for more information. The default is 'reduced', and to
maintain backward compatibility with earlier versions of numpy both
it and the old default 'full' can be omitted. Note that array h
returned in 'raw' mode is transposed for calling Fortran. The
'economic' mode is deprecated. The modes 'full' and 'economic' may
be passed using only the first letter for backwards compatibility,
but all others must be spelled out. See the Notes for more
explanation.
Returns
-------
q : ndarray of float or complex, optional
A matrix with orthonormal columns. When mode = 'complete' the
result is an orthogonal/unitary matrix depending on whether or not
a is real/complex. The determinant may be either +/- 1 in that
case.
r : ndarray of float or complex, optional
The upper-triangular matrix.
(h, tau) : ndarrays of np.double or np.cdouble, optional
The array h contains the Householder reflectors that generate q
along with r. The tau array contains scaling factors for the
reflectors. In the deprecated 'economic' mode only h is returned.
Raises
------
LinAlgError
If factoring fails.
Notes
-----
This is an interface to the LAPACK routines dgeqrf, zgeqrf,
dorgqr, and zungqr.
For more information on the qr factorization, see for example:
http://en.wikipedia.org/wiki/QR_factorization
Subclasses of `ndarray` are preserved except for the 'raw' mode. So if
`a` is of type `matrix`, all the return values will be matrices too.
New 'reduced', 'complete', and 'raw' options for mode were added in
NumPy 1.8.0 and the old option 'full' was made an alias of 'reduced'. In
addition the options 'full' and 'economic' were deprecated. Because
'full' was the previous default and 'reduced' is the new default,
backward compatibility can be maintained by letting `mode` default.
The 'raw' option was added so that LAPACK routines that can multiply
arrays by q using the Householder reflectors can be used. Note that in
this case the returned arrays are of type np.double or np.cdouble and
the h array is transposed to be FORTRAN compatible. No routines using
the 'raw' return are currently exposed by numpy, but some are available
in lapack_lite and just await the necessary work.
Examples
--------
>>> a = np.random.randn(9, 6)
>>> q, r = np.linalg.qr(a)
>>> np.allclose(a, np.dot(q, r)) # a does equal qr
True
>>> r2 = np.linalg.qr(a, mode='r')
>>> r3 = np.linalg.qr(a, mode='economic')
>>> np.allclose(r, r2) # mode='r' returns the same r as mode='full'
True
>>> # But only triu parts are guaranteed equal when mode='economic'
>>> np.allclose(r, np.triu(r3[:6,:6], k=0))
True
Example illustrating a common use of `qr`: solving of least squares
problems
What are the least-squares-best `m` and `y0` in ``y = y0 + mx`` for
the following data: {(0,1), (1,0), (1,2), (2,1)}. (Graph the points
and you'll see that it should be y0 = 0, m = 1.) The answer is provided
by solving the over-determined matrix equation ``Ax = b``, where::
A = array([[0, 1], [1, 1], [1, 1], [2, 1]])
x = array([[y0], [m]])
b = array([[1], [0], [2], [1]])
If A = qr such that q is orthonormal (which is always possible via
Gram-Schmidt), then ``x = inv(r) * (q.T) * b``. (In numpy practice,
however, we simply use `lstsq`.)
>>> A = np.array([[0, 1], [1, 1], [1, 1], [2, 1]])
>>> A
array([[0, 1],
[1, 1],
[1, 1],
[2, 1]])
>>> b = np.array([1, 0, 2, 1])
>>> q, r = LA.qr(A)
>>> p = np.dot(q.T, b)
>>> np.dot(LA.inv(r), p)
array([ 1.1e-16, 1.0e+00])
Lab: Least-squares regression various ways¶
Load Boston house prices dataset.
Formulate linear system and try using inverse and pseudoinverse to solve.
from sklearn.datasets import load_boston
boston = load_boston()
print(dir(boston))
print(boston.data.shape)
print(boston.feature_names)
print(boston.DESCR)
['DESCR', 'data', 'feature_names', 'filename', 'target']
(506, 13)
['CRIM' 'ZN' 'INDUS' 'CHAS' 'NOX' 'RM' 'AGE' 'DIS' 'RAD' 'TAX' 'PTRATIO'
'B' 'LSTAT']
.. _boston_dataset:
Boston house prices dataset
---------------------------
**Data Set Characteristics:**
:Number of Instances: 506
:Number of Attributes: 13 numeric/categorical predictive. Median Value (attribute 14) is usually the target.
:Attribute Information (in order):
- CRIM per capita crime rate by town
- ZN proportion of residential land zoned for lots over 25,000 sq.ft.
- INDUS proportion of non-retail business acres per town
- CHAS Charles River dummy variable (= 1 if tract bounds river; 0 otherwise)
- NOX nitric oxides concentration (parts per 10 million)
- RM average number of rooms per dwelling
- AGE proportion of owner-occupied units built prior to 1940
- DIS weighted distances to five Boston employment centres
- RAD index of accessibility to radial highways
- TAX full-value property-tax rate per $10,000
- PTRATIO pupil-teacher ratio by town
- B 1000(Bk - 0.63)^2 where Bk is the proportion of blacks by town
- LSTAT % lower status of the population
- MEDV Median value of owner-occupied homes in $1000's
:Missing Attribute Values: None
:Creator: Harrison, D. and Rubinfeld, D.L.
This is a copy of UCI ML housing dataset.
https://archive.ics.uci.edu/ml/machine-learning-databases/housing/
This dataset was taken from the StatLib library which is maintained at Carnegie Mellon University.
The Boston house-price data of Harrison, D. and Rubinfeld, D.L. 'Hedonic
prices and the demand for clean air', J. Environ. Economics & Management,
vol.5, 81-102, 1978. Used in Belsley, Kuh & Welsch, 'Regression diagnostics
...', Wiley, 1980. N.B. Various transformations are used in the table on
pages 244-261 of the latter.
The Boston house-price data has been used in many machine learning papers that address regression
problems.
.. topic:: References
- Belsley, Kuh & Welsch, 'Regression diagnostics: Identifying Influential Data and Sources of Collinearity', Wiley, 1980. 244-261.
- Quinlan,R. (1993). Combining Instance-Based and Model-Based Learning. In Proceedings on the Tenth International Conference of Machine Learning, 236-243, University of Massachusetts, Amherst. Morgan Kaufmann.
Nonlinear regression¶
(easy way) - Linear regression after nonlinear transformation
Old: \begin{align} \mathbf y = \beta_0 + \beta_1 \mathbf x_{(1)} + \beta_2 \mathbf x_{(2)} +...+ \beta_n \mathbf x_{(n)} + \varepsilon , \; \; \; \mathbf x_{(i)} = \begin{bmatrix} x_{(i),1} \\x_{(i),2} \\ \vdots \\ x_{(i),m} \end{bmatrix} \end{align}
New: \begin{align} \mathbf y = \beta_0 + \beta_1 \mathbf x'_{(1)} + \beta_2 \mathbf x'_{(2)} +...+ \beta_n \mathbf x'_{(n)} + \varepsilon, \;\;\; \mathbf x'_{(i)} = \begin{bmatrix} f_{(i)}(x_1) \\f_{(i)}(x_2) \\ \vdots \\ f_{(i)}(x_m) \end{bmatrix} \end{align}
...Feature Engineering
Polynomial regression - simple powers of data¶
New model: $\mathbf y = \beta_0 + \beta_1 \mathbf x'_{(1)} + \beta_2 \mathbf x'_{(2)} +...+ \beta_n \mathbf x'_{(n)} + \varepsilon$
$$\mathbf x'_{(i)} = \begin{bmatrix} f_{(i)}(x_1) \\f_{(i)}(x_2) \\ \vdots \\ f_{(i)}(x_m) \end{bmatrix} = \begin{bmatrix} x_{1}^i \\x_{2}^i \\ \vdots \\ x_{m}^i \end{bmatrix}$$Exercise: write the model equations out for 3D case, i.e. $\bf x$ is just $x_1$, $x_2$, and $x_3$.
Polynomial regression¶
show_polyfit_ho_example_results(dosage,conc_noisy,(1,2,3,4,5,6));
order: 1, residual: 56.6204627002199 order: 2, residual: 43.28483436298878 order: 3, residual: 38.90189854595423 order: 4, residual: 21.166171682768383 order: 5, residual: 13.416800916508517 order: 6, residual: 1.8203750586044797e-11