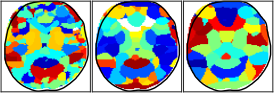
Brain parcellation is important for exploiting neuroimaging data. Variability in physiology between individuals has led to the need for data-driven approaches to parcellation, with recent research focusing on simultaneously estimating and partitioning the network structure of the brain. We view data preprocessing, parcellation, and parcel validation from the perspective of predictive modeling. The goal is to identify parcels in a way that best generalizes to unseen data. We utilize an uncertainty quantification approach from image science to define parcels as groups of unresolvable variables in the predictive model.