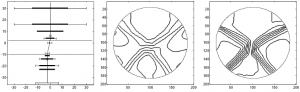
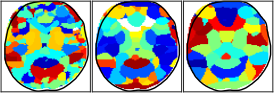
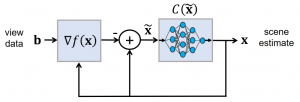
Model-based machine learning methods incorporate domain knowledge from the physical forward model of an inverse problem to reduce the need for training data. In this research, we show how this can be used to address challenging limitations such as occlusion. We combine a convolutional neural network with a novel computational reconstruction method that combines source and attenuation distributions in order to model occlusion.
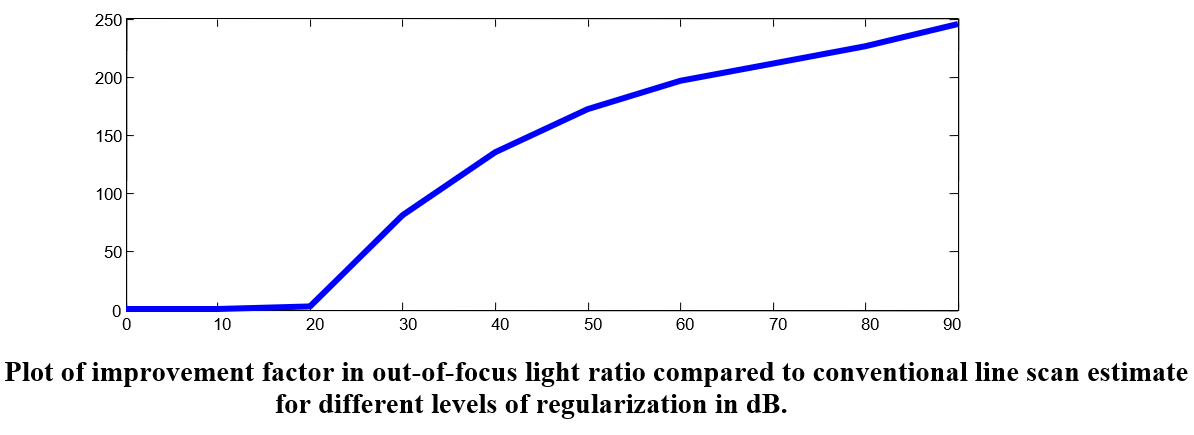
We demonstrate the ability to quickly learn to address reconstruction artifacts and opacity, forming a significantly improved final image of the scene based on as little as a single training image. The algorithm can be implemented efficiently and scaled to large problem sizes.
Link: Model-based machine learning for computational reconstruction of opacity and missing information